Warmer seas linked to disease epidemics affecting Pacific starfish and Atlantic lobster
|
A study carried out just off of the coast of Washington State in America observed the rising of temperature in the Atlantic and Pacific oceans which is linked to disease epidemic. Epidemic is a widespread occurrence of an infectious disease in a community at a particular time. Different disease outbreaks were seen in the Starfish off the west coast of the US and Canada and the Lobster off the coast of New England. The starfish endured high levels of a physically degenerative wasting disease which cause them to loose their limbs and die between 2013 and 2014. Whereas the lobsters experienced an increase in shell disease. Scientists conducting this study have linked both of these diseases to the rising temperature of the sea. Drew Harvell, Cornell professor of Ecology and evolutionary biology and senior author of this study said that "This outbreak came so quickly, by the time we knew enough about it, a lot of it had already happened". He says that this year they are looking out especially in areas in Alaska where there used to be copious amounts of Sunflower starfish and are now gone. They suppose it could have been a virus that knocked these starfish out of the ocean which becomes more active and dangerous in warmer waters. This outbreak occurred when the water temperature started to rise. In lab conditions they put some starfish into cold water and some into warm water and found that the ones in warmer water succumb to disease more quickly. They also found that the disease was more prevalent in temperatures between 12.2 and 18.8 degrees celsius. In the lab they also tested the disease that affected the lobsters and found that at colder temperatures the disease made less progress then at warmer temperatures. And as it got warmer, it affected females more then males. This may be because female lobsters shed their shells less frequently then male lobsters. Professor Harvell says that they can predict when disease will outbreak in these species based on temperature projections of water.
Warmer seas linked to disease epidemics affecting Pacific starfish and Atlantic lobster
0 Comments
The human genome project is a very important experiment in terms of advancing science and furthering research. It was a project with the aim to find the base sequence of the entire human genome. This project was completed worldwide and although, after 13 years, the entire sequence was in fact found, there are some power struggles with it. For starters the genome was exactly that. It was A human genome rather then THE human genome. But despite this there are positive outcomes of this, helping the advancement of science. The project, for starters, drove rapid improvements in base sequencing techniques which allowed a draft sequence to be published much sooner then expected in 2000. The complete sequence was published in 2003. One discovery during this project was that it is possible to predict which base sequences are protein-coding genes. There are approximately 23,000 in the human genome. Another discovery was that most of the genome is intact not transcribed. This is known commonly as "junk DNA". But within these regions there are elements that affect gene expression as well as highly repetitive sequences, called satellite DNA.
Work today continues to find variations in sequence between different individuals. The vast majority of base sequences are shared by all humans giving us genetic unity but there are also many single nucleotide polymorphisms which contribute to human diversity. Since the publication of this project the base sequence of many other species has been determines. Comparisons between these genomes reveal aspects of the evolutionary history of living organisms that were previously unknown.
causes hemoglobin molecules to stick together in tissues with low oxygen concentrations. The bundles of hemoglobin molecules that are formed are rigid enough to distort the red blood cells into a sickle shape. These sickle cells cause damage to tissues by becoming trapped in blood capillaries, blocking them and reducing blood flow. When sickle cells return to their normal shape. These changes occur time after time, as the red blood cells circulate. Both the hemoglobin and the plasma membrane are damaged and the life of a red blood cell can be shortened to as little as 4 days. The body cannot replace red blood cells at a rapid enough rate and anaemia therefore develops.
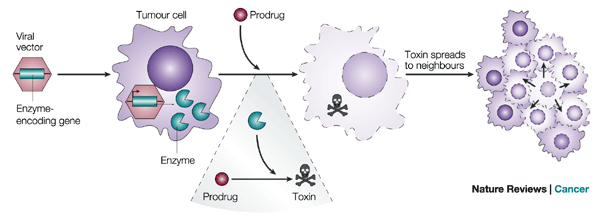
Today in class we did an experiment where we used thin layer chromatography. Procedure: 1) Add a small spatula measure of anhydrous sodium sulphate(VI) to the bottom of a plastic eppendorf tube. This is to remove any moisture that may be present. 2) Place small pieces of a spinach in the bottom of the eppendorf tube. Add 5 drops of Solvent A. consisting of 2 parts ethyl ethanoate and 3 parts proponent. 3) Press/stir the spinach with the end of forceps to extract as much colour as possible. 4) Use a fine brush to place a small dot of the coloured solvent about 5mm from the bottom edge of the TLC plate. Mark this position on the side of the plate with a pencil. 5) Gently blow on the dot to dry it before pitting another dot of coloured solution on the top of it. Continue to do this until the dot is not to faint but not to strong. 6) Use a dropper pipette to place o.5cm^3 of Solvent B into a clean vial so that the depth is about 5mm. The level of the solvent must be below the level of the dot on the chromatography plate. 7) Put the TLC plate into the vial and replace the screw-top lid. 8) After a few minutes, when the 'solvent front' has moved 7/8ths of the way up the plate, remove the plate from the vial using forceps and let it dry. Mark the position of the solvent front. What the results mean: In order of line appearance from the top end of the TLC plate, where the solvent front is. - First seen is the B-catorene - Second is Pheophytin A - Then is Pheophytin B - Followed by the Chlorophyll A - After is Chlorophyll B - Then Lutein - And lastly follows any other Xanthophyll's BBC News published an article on the 8th of December about a new gene therapy that can kill off prostate cancer cells. The therapy technique is able to modify the cancer cells so that the body can attack and kill them. It causes the cells to 'self-destruct' therefore gaining it's name 'suicide gene therapy'. It involves genetically modifying the cancer cells to signal the body's immune system to attack them. They do this because the body doesn't usually recognise cancer cells as the enemy as they are evolved from normal healthy cells. A virus is used to carry the gene therapy to the tumour cells and it results in the cells self-destructing, telling the body's immune system to launch an attack. This research project was led by US scientist at Huston Methodist Hospital, Texas, and they showed that this therapy increased the survival rate in patients by 20% but say that more research is needed to justify this and see the full effectiveness of this treatment. But also say that when combined with radiotherapy, suicide gene therapy is a very promising treatment for prostate cancer in the future.
BBC News "'Suicide' gene therapy kills prostate cancer cells" Article The below image shows the delivery of the suicide gene and its by standing effects: In todays class we did practical work which involved taking milk and transforming it into cat milk, milk without lactose.
Method: 1) Mix 2cm^3 of lactase enzyme with 8cm^3 of 2% sodium alginate solution 2) Add this mix to 1.5% calcium chloride solution a drop at a time, this created gel like balls. 3) Use a tea strainer to separate the balls from the calcium chloride solution 4) Pack a small piece of gauze and the balls into a syringe with an attached tube of plastic going into a beaker with a tap attached to maintain control over the milk flow. 5) Then pour the milk through the balls into the beaker and test the milk for lactose levels every time around. Unfortunately we didn't have time to make it very lactose free in our class but we our results showed us was that there was less and less lactose in the milk each time. In class we did an experiment to get rid of the amylose in a solution using the enzymes in our saliva.
Scientists from the University of Edinburgh and the University of Dundee in the United Kingdom have found a way in which eating ice cream can be less sticky in the heated weather of summer by making it melt less quickly. They discovered a protein called BsIA that is usually found in large bacterial communities in structures called biofilm which can be used to keep everything combined in ice cream. It therefore changes some of the properties of ice cream like making it smoother and more resistant to melting. Emulsifiers, small, fat-like molecules that keep oil and water mixed, are used to make ice cream but these scientist replace that with the BsIA protein. The protein creates a hydrophobic coat on the outer surface of the biofilm and the way it's created allows for a stable interaction between substances that usually repel each other like oil and water. In ice cream the important interactions are between oil and water, and air bubbles and water. Therefore making this protein the perfect substitute in ice cream and having the added effect of making it melt less quickly making it more enjoyable for us to eat!
Live science 'No More Sticky Mess! Scientists Develop Slower-Melting Ice Cream' Article The Mirror's 'Sick of your ice-cream letting too quickly? Scientists might have the answer' Article In today's society, psychologists all over the world wonder about memories and just how exactly they are formed. In Seoul, Korea Scientist Jun Cho and his team used ribosome profiling and RNA-sequencing to look in more detail at the role of genes in memory formation. He used mice, after they had gone through fear conditioning, and compared then to mice without fear condition and found interesting results. The fear conditioning consisted of the mice experiencing small electric shocks in a specific environment so that they form a strong, negative memory with the environment. The scientist then analysed the hippocampi of the conditioned mice and those who weren't conditioned at different intervals after conditioning. His research looked for genes that were being expressed differently in the conditioned mice and the non-conditioned mice and identified 104 genes. Around half of those were being regulated by oestrogen receptor alpha (ESR1) . Further research following this found that the ESR1 impaired memory formation in mice during the hippocampus-depentant tasks they were put through when being investigated. This essentially suggests that the ESR1 may play an important role in modulating gene-regulatory networks after learning.
Gene suppression helps form memories - Science daily ESR1: Embryonic Stem Cells
|
AuthorI am Abigail Ramsey, A student at Ardingly College. I am studying the IB and one of my subjects is high level biology. This is a blog of interesting things i find on the internet, cool labs we do in class or anything else related to biology. Archives
October 2016
|